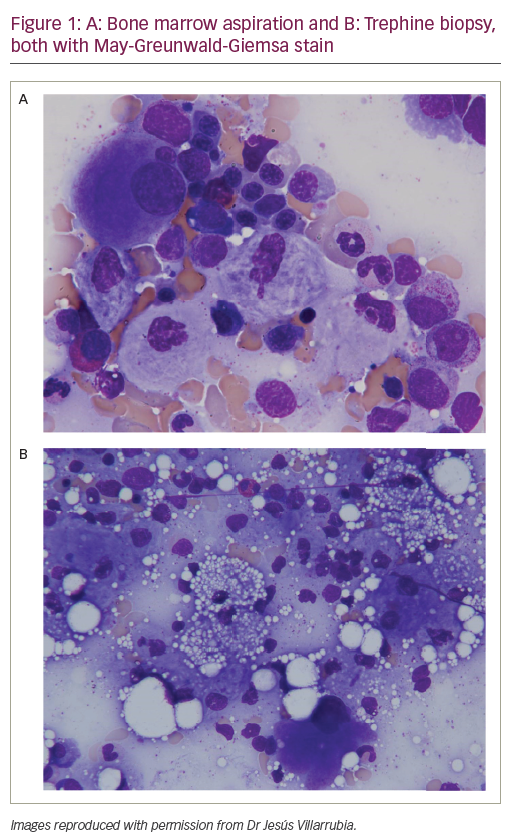
Fanconi anaemia (FA) is a genetic, life-threatening disorder1,2 featuring progressive bone marrow failure, birth defects, leukaemia, increased incidence of solid tumours, spontaneous chromosomal instability and hypersensitivity to cross-linking reagents.1,2 Because of FA’s relationship to DNA damage hypersensitivity and its association with susceptibility to neoplastic transformation, the FA research field has gained much attention in recent decades, especially after the discovery of the connection between FA and the well-known breast cancer (BRCA) genes.
Fanconi anaemia (FA) is a genetic, life-threatening disorder1,2 featuring progressive bone marrow failure, birth defects, leukaemia, increased incidence of solid tumours, spontaneous chromosomal instability and hypersensitivity to cross-linking reagents.1,2 Because of FA’s relationship to DNA damage hypersensitivity and its association with susceptibility to neoplastic transformation, the FA research field has gained much attention in recent decades, especially after the discovery of the connection between FA and the well-known breast cancer (BRCA) genes.
Clinical Features
FA is a very heterogeneous condition clinically, and about 70% of patients display a wide variety of abnormalities, classic among which are missing or misshapen thumbs and radii, kidney, gastrointestine and neurocognitive and developmental delay.3–10 The most important clinical feature of FA is marrow failure in the form of aplastic anaemia, myelodysplastic syndrome or acute myeloid leukaemia (AML).3,4 The number of FA children surviving childhood has greatly increased with the advent of improved outcomes after bone marrow transplantation for bone marrow failure and AML, but those patients surviving until adulthood have to face a greatly increased risk of solid tumours, typically squamous cell carcinomas such as head and neck and gynaecological tumours.11,12
Because FANCD1/BRCA2 conveys an inherited risk of breast, ovarian and pancreatic cancer for individuals carrying a single mutated allele, the FA pathway is thought to be involved in cancer development. Many studies support the notion that somatic inactivation of the FA pathway may contribute to the pathogenesis of sporadic cancers, including AML.13 For pancreatic cancer, although screens for germline mutations in FANCC, FANCA and FANCG failed to detect any pathogenic alleles in familial tumours,14 truncations of FANCC were found in sporadic pancreatic cancers of early-onset cases.15 Expression of FANCF is decreased in most ovarian cancers compared with those in normal ovarian tissue,16 in part because of methylation of the FANCF promoter, suggesting a role for epigenetic modifications in FA tumorigenesis.16 Restoration of this pathway is associated with demethylation of FANCF, leading to acquired cisplatin resistance.6–8
Fanconi Anaemia Genes
At the cellular level, the distinguishing and diagnostic features of FA are spontaneous genomic instability and hypersensitivity to DNA interstrand cross-linkers such as mitomycin C (MMC) and diepoxybutane (DEB). FA complementation groups were established by the pair-wise fusion of patient-derived cell lines and the subsequent resistance to genotoxic reagents. The genes were subsequently cloned via expression, purification, positional and candidate approaches.17–28,29 A summary of identified FA genes is shown in Table 1.
Protein Biochemistry of Fanconi Anaemia
Core Complex Formation
Eight FA genes (FANCA, FANCC, FANCB, FANCG, FANCF, FANCE, FANCL and FANCM) encode for members of a multi-subunit protein complex termed the FA core complex. Although the stoichiometry is unclear, most subunits are required for general complex stability and proper nuclear localisation, which is essential for mono-ubiquitilation of the downstream effector FANCD2.17 FANCA and FANCG readily associate and stabilise each other.30 Other known interacting pairs within the FA core complex include FANCE/FANCC31 and FANCB/FANCL.32 FANCF functions as a molecular adaptor, bridging between the subcomplexes A:G and C:E,33 whereas FANCM may link the subcomplex B:L to the subcomplex A:G.32 The FA core complex is also reported to weakly associate with other DNA repair proteins, including Bloom’s helicase,34 topoisomerase IIIα35 and replication protein A.35 Although the functional significance of these inter-complex associations is uncertain, they nevertheless imply a role of FA in DNA damage response. The amount of the chromatin-bound fraction of FA complex increases in the S phase when the FA pathway is activated,36 and is facilitated by FANCM.24 Similar results were observed in a newly developed cell-free Xenopus system.37 The loading and unloading of the xFA core complex is strictly dependent on entry and exit during the S phase, supporting the role of FA in DNA replication.
Mono-ubiquitilation of FANCD2
FANCD2 is mono-ubiquitilated on lysine 561 in response to DNA damage38 as well as S-phase progression,39 which is dependent on the FA core complex. This is a pivotal event to activate the FA pathway. FA-D2 cell lines expressing mutant FANCD2 (K561R) are as sensitive to MMC as parental cell lines. The whole FA core complex is hypothesised to be the ubiquitin E3 ligase for FANCD2 with FANCL as the catalytic subunit. FANCE may serve as a link between FANCD2 and the core complex, as it is found to be bound to both.31 In addition, knockdown of the E2 ligase UBE-2T, a binding partner of FANCL, leads to a drastic reduction of the level of mono-ubiquitilated FANC-D2 (FANCD2-L isoform).40 However, the identity of FANCD2-specific E3 ligase remains a question, because FANCL has not been demonstrated to mono-ubiquitilate FANCD2 in vitro. In addition, the core complex clearly has additional activities. A chicken model of FA has demonstrated that a ub-tailed version of FANCD2 can correct chicken FA-D2 mutants but not core complex mutants. Following mono-ubiquitilation, FANCD2 is recruited to chromatin foci, where it is co-localised with other DNA repair proteins, including BRCA1 and RAD51.15,17 Knockout experiments in chicken DT40 cells demonstrate that the FA core complex is also necessary for the chromatin association of FANCD2-L isoform.41 After completion of the DNA repair response, the deubiquitilating enzyme USP1 catalyses the removal of the FANCD2 ubiquitin moeity42 to downregulate the expression of mono-ubiquitilated D2. Interestingly, the newly identified FANCI is a paralogue of FANCD2 and is mono-ubiquitilated in a similar fashion. FANCI and FANCD2 assemble to form a so-called I-D complex, and mono-ubiquitilation of either one is required for the maintenance of mono-ubiquitilation of the other (see Figure 1).
Fanconi Anaemia and DNA Repair Pathways
In mammalian cells, the nucleotide excision repair pathway is responsible for the first step of interstrand cross-link (ICL) incision and is not dependent on the FA pathway.43–46 Following incision, ICLs are processed into double-strand breaks (DSBs) upon replication fork progression,45,47 triggering orchestrated repair responses, including activation of the FA pathway and restarting the replication forks. The mechanistic details are largely unknown; nonetheless, there is genetic evidence that FA proteins work upstream of two DNA repair pathways, translesion synthesis (TLS) and homologous recombination repair (HRR), presumably by stabilising the stalled replication forks from collapse. DT40 cells with defects in TLS (rev1 and rev3) and HRR (rad51 and brca2) both show FA-like phenotypes such as hypersensitivity to MMC and decreased sister chromatid exchange rate.43 Analysis of the spontaneous mutation rate also demonstrates that FA proteins promote both error-free and error-prone repairs,48,49 suggesting involvement of both HRR and TLS in ICL repair. This idea is further strengthened by physical interactions between FANCG, FANCD2 and FANCD1/BRCA2,50,51 presumably to enhance the nucleation of RAD51 filaments. In chicken DT40 cell lines, FANCD2 co-localises with REV1 specifically following replication arrest but not X-ray,48,49 which further supports this idea.
Fanconi Anaemia and Signal Transduction
Fanconi Anaemia and DNA Damage Checkpoint Kinase ATR
In response to cross-links detected during DNA replication, the cell-cycle checkpoint kinase ATR is essential for phosphorylation of FANCD2, which is probably a prerequisite for FANCD2 mono-ubiquitilation. NBS1-FANC-FANCD2 and CHK1 are two independent branches downstream of ATR in order to mediate an intra-S-phase checkpoint.52 NBS1, the protein defective in Nijmegen breakage syndrome (NBS), is part of the nuclear MRN complex (MRE11/RAD50/NBS1) involved in the processing of DSBs.53 An intact FA core complex is required for phosphorylation of NBS1 on serine 343 and relocalisation of MRN complex to FANCD2 nuclear foci after DNA damage.54 Conversely, phosphorylated NBS1 participates in phosphorylation of FANCD2, permitting multiple levels of regulation of the intra-S-phase checkpoint arrest. ATR also directly phosphorylates FANCD2 on threonine 691 and serine 717, which is also required for the establishment of an intra- S-phase checkpoint.55,56 CHK1 kinase is known to mediate the post-replication checkpoint and has a critical role in regulating chromosome stability.57 Inhibition of CHK1 leads to loss of mono-ubiquitilated FANCD2 and hypersensitivity to DNA cross-linkers, suggesting CHK1 couples ATR signalling to activate the FA pathway. ATR also phosphorylates histone H2AX on serine 139 at stalled replication forks, which in turn recruits mono-ubiquitilated FANCD2 in a BRCA1-dependent manner. Inhibition of phosphorylation of H2AX renders cells to spontaneous chromosome aberrations and hypersensitivity to MMC, as with FA mutant cells.58
Phosphorylation of Fanconi Anaemia Core Complex Proteins
A number of FA member proteins are phosphorylated in response to DNA damage or cell-cycle progression. FANCM contains multiple predicted ATR phosphorylation sites and is hyper-phosphorylated following MMC treatment.24 FANCA is phosphorylated in an FA-complex-dependent manner,59 and is also reported to associate with the IkappaB kinase (IKK) signalling via interaction with IKK2, a kinase required for rapid, stress-dependent phosphorylation of the FA core complex.60 FANCG is phosphorylated on serine-7, a modification required for full correction of MMC resistance in FA-G cells,61 as well as at serine 383 and serine 387 during mitosis,62 presumably to regulate the exit of FA complexes from the nucleus. The FANCE subunit of the core complex is phosphorylated on two conserved sites (threonine 346 and serine 374) by CHK1.63 Interestingly, the non-phosphorylatable FANCE mutant (FANCE-T346A/S374A) supports FANCD2 mono-ubiquitilation, FANCD2 foci assembly and normal S-phase progression, but fails to complement the mitomycin C hypersensitivity of the transfected FA-E mutant cells, suggesting yet another role for CHK1.63
Non-nuclear Roles of the Fanconi Anaemia Pathway
Tumour Necrosis Factor-α, Interferon-γ and Pro-apoptotic State
Tumour necrosis factor-α (TNF-α) and interferon-γ (IFN-γ) are well-known negative modulators of haematopoiesis by triggering excessive apoptosis. In vivo, TNF-α and IFN-γ are overexpressed in stimulated bone marrow lymphocytes from FA-C patients.64 In vitro, both the number and size of erythroid colony-forming units and erythroid burst-forming units grown from FA patients are subject to inhibition by TNF-α.64 Murine models further support the theory that FA proteins are involved in cytokine signalling pathways and that cytokine-mediated apoptosis has a major role in the pathogenesis of bone marrow failure observed in FA patients.65
Increased apoptosis may be mediated via TNF-related apoptosis-inducing ligand in FA-A cells, although FA-C cells are resistant to TRAIL.66,67 Protein kinase regulated by RNA (PKR) plays a critical role in cell growth and apoptosis68 and is implicated in the FA pathway. In primary human bone marrow cells, mutation in FANCA, FANCC or FANCG markedly increases the association between PKR and FANCC, leading to hyperactivation of PKR and hypersensitivity to cytokine-mediated cytotoxicity.69,70 FANCC binds to the signal transducer and activator of transcription 1 (STAT1) following IFN-γ treatment and is required for proper docking of STAT1 at the IFN-γ receptor α-chain71 for survival cues. Together, augmented activity of PKR and failed activation of STAT1 by IFN-γ may thus tip the balance from survival to death in FA-C haematopoietic progenitor cells.
Fanconi Anaemia Oxygen Sensitivity
There is an extensive body of evidence indicating that the FA pathway has a role in oxidative stress response. FANCC associates with proteins with redox activities such as NADPH cytochrome P450 reductase72 and glutathione S-transferase.73 FANCG protein interacts with cytochrome P450 2E1, a P450 protein known to be involved in redox biotransformation of xenobiotics, including MMC.74 Microarray experiments show consistent overexpression of proteins involved in oxidative metabolism, including nuclear factor 1, heat shock protein 70kDa and cyclo-oxyenase 2 in FA mutant cells.75,76 The role of the FA pathway in oxidative stress response is further supported by data from primary and immortalised cell cultures, indicating a pro-oxidant state in FA cells.77,78 More significantly, chromosome abnormalities,79–81 cell-cycle defects82 and pro-apoptotic propensity83,84 in FA cells can be corrected by antioxidant enzymes or low-molecular- weight antioxidants (such as thioredoxin) or hypoxia.85 A recent study found that hypoxia–re-oxygenation cycles mimicking the condition faced by migrating haematopoietic cells induce premature senescence arrest and inhibit expansion of fancc-/- BM cells, suggesting a novel mechanism for bone marrow failure in FA.86 ■