The relapse rate among patients with early-stage non-small-cell lung cancer (NSCLC) is 40% within five years after potentially curative treatment.1 The current staging system for NSCLC is inadequate for predicting the outcome of treatment. To guide clinical decisions on the optimum treatment regimen, there is clearly a need to accurately identify patients at high risk for recurrent or metastatic disease. Adenocarcinomas account for 40% of NSCLC and they have shown an increasing incidence, particularly in females and non-smokers.2 Adenocarcinomas represent a heterogenous group of tumors and are known to exhibit the highest degree of morphological and clinical diversitiy, which reflects patient survival.3 This review highlights the molecular characteristics that may influence metastatic behavior in lung adenocarcinoma.
Molecular Signatures
The ability of cells to gain metastatic potential is an intrinsic property of the primary tumor, which is substantiated by the high correlations between clinical outcome and gene expression profiles of a variety of primary tumors. Gene expression profiling with the use of microarrays or polymerase chain reaction (PCR) has been shown to estimate the prognosis for patients with lung cancer. Moreover, based on recent investigations into the nature of adenocarcinoma of the lung, it has become obvious that several distinct molecular diseases are lumped together under this morphological entity. More than one molecular signature of lung adenocarcinoma associated with metastasis and survival have been reported.4–14 Chen et al. identified a five-gene signature that was significantly associated with relapse-free and overall survival among patients with NSCLC.4 Also, Potti et al. developed a lung metagene model capable of predicting recurrence for individual patients significantly better than clinical prognostic factors and which was consistent with early stages of NSCLC.5
Patients with a high-risk gene signature could benefit from adjuvant therapy, whereas those with low risk could be spared unnecessary treatment. A combined squamous cell and adenocarcinoma molecular classifier was able to predict significant differences in overall survival and thus to capture the majority of NSCLC.6 Recently, Beer et al. reported that microarray analysis of gene expression in lung adenocarcinomas can also predict outcome in stage I patients.7 Through comparison of gene expression in primary adenocarcinoma samples and the same expression profile in metastatic tumors derived from unmatched primary adenocarcinomas, Ramaswamy et al. identified a metastasis-associated gene expression signature significantly correlated with survival.8 By using a 318-gene set classifier in primary lung adenocarcinomas, Xi et al. were able to predict lymph node metastasis and overall outcome.9 In their model, Guos et al. identified a 37-gene signature not only for the detection of specific high-risk lung adenocarcinoma patients, but also for the prediction of the stage as well as the differentiation of the tumors.10 The gene ontology composition of the above signatures together with other studies11–14 comprises genes that have biological relevance to disease recurrence, such as angiogenesis, apoptosis, cell cycle, and cell motility and adhesion. The identification of these genes may reveal targets for the development of therapies for lung cancer and explores the kind of response or resistance to standard or novel therapies. Such research suggests that molecular signatures can guide the choice of treatment for primary and adjuvant therapy.
MicroRNAs
MicroRNAs are a class of small non-protein-coding RNAs that can act as endogenous RNA interference. MicroRNAs can post-transcriptionally regulate the expression of hundreds of their targeted genes, thereby controlling a wide range of biological functions such as cellular proliferation, differentiation, and apoptosis.15 They have been found to be abnormally expressed in several types of cancer, suggesting that miRNAs play a substantial role in the pathogenesis of human cancers. MicroRNA microarray analysis has identified statistically unique profiles correlated with survival of lung adenocarcinomas, including those classified as stage I.16 Two of the eight miRNA signatures, high expression of hsa-mir-155 and low expression of hsa-let-7a-2, correlated with poor survival of lung adenocarcinomas. Recently, it was shown that downregulation of the let-7 family could result in the upregulation of RAS and induce oncogenesis in human lung cancer, and is associated with reduced survival in NSCLC.17,18 These results were confirmed recently when realtime reverse-transcription (RT)-PCR was used to discover a five-miRNA signature.19 This method could be more clinically applicable since it can use a small amount of bronchoscopy sampling or paraffin-embeded specimens. However, further investigation is needed to identify the targets of the miRNAs and to experimentally correlate them with metastatic behavior in lung carcinogenesis.
Epidermal Growth Factor Receptor
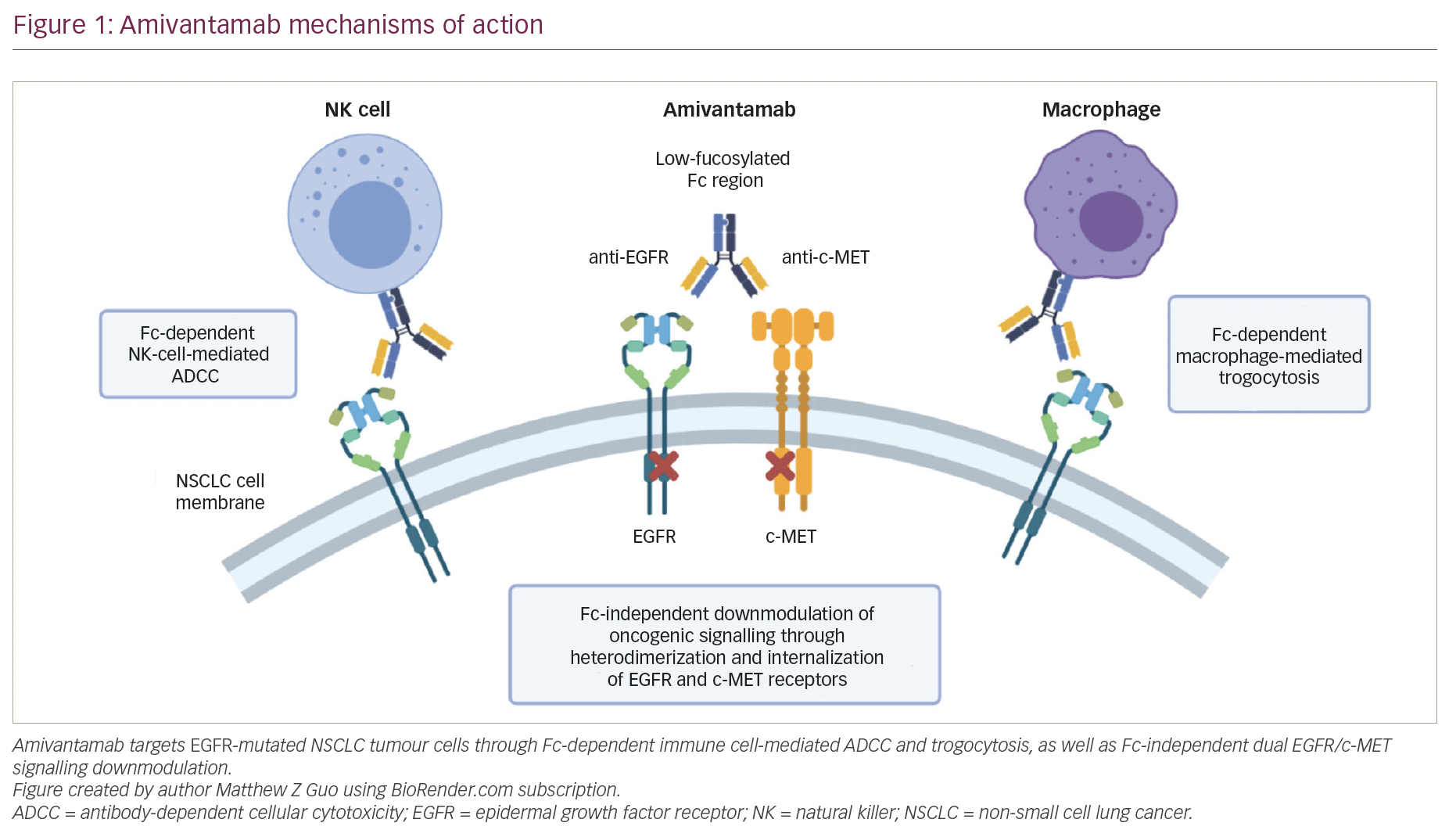
Epidermal growth factor receptor (EGFR) is a receptor protein tyrosine kinase (RTK) deeply involved in the carcinogenesis of NSCLC, mainly adenocarcinomas.20 The EGFR induces cancer via at least three major mechanisms: overexpression of EGFR ligands, amplification of EGFR, and activating mutations in the EGFR gene.21 Recent meta-analyses22–25 showed that EGFR mutations, especially deletions in exon 19 and L858R point mutations in exon 21, are more frequent in East-Asian ethnicity, neversmokers, adenocarcinoma histology, and female gender. Increased copy number of EGFR, as assessed by fluorescence in situ hybridization (FISH) assay, was associated with lymph node metastasis, more advanced pathological stage, and poor prognosis. This alteration has been described in NSCLC, especially adenocarcinomas, and may occur in mutant or wildtype tumors.26,27 A recent study revealed that increased copy number of the EGFR gene seems to be a progression event independent of the initial alterations, being more than four times more frequent in invasive lesions than in non-invasive lesions.28 Lung adenocarcinomas that harbor somatic mutations in the EGFR are highly likely to respond to the EGFR RTK inhibitors gefitinib and erlotinib.29 One possible explanation for this phenomenon is that the cancer cells are ‘addicted’ to signaling via the mutant EGFRs and die when the mutant oncoprotein is inactivated. Available data on the prognostic value of EGFR mutations are conflicting. The molecular analysis of patients treated in the TRIBUTE (erlotinib) and INTACT (gefinitib) trials suggested a prolonged survival in patients with mutations;29,30 no significant differences were found in patients undergoing resections of NSCLC31 or lung adenocarcinomas.11 In contrast, there is no doubt about the predictive value of EGFR mutations in tumor response, which ranged between 65 and 92% in patients with mutations and was 9–13% in patients without mutations.32
RAS
The RAS proteins are pivotal regulators of cellular proliferation, differentiation, motility, and apoptosis, with mutations in KRAS occurring in 30–50% of lung adenocarcinomas, predominantly in smokers.25 In total, 80% of KRAS mutations occur in codon 12, occasionally in codon 13, and rarely in codon 61.33 The negative prognostic impact of KRAS mutations has been demonstrated in several studies and confirmed in a recent metaanalysis. 33 Mutations of KRAS may be predictive of resistance to chemotherapy. Adjuvant chemotherapy did not confer survival advantage in patients whose tumors had RAS mutations.34 The correlation of KRAS status to clinical outcomes in the TRIBUTE study showed that patients with KRASmutant tumors not only fail to benefit from erlotinib plus chemotherapy, but may experience decreased survival and time to progression (TTP).35 Despite demonstrating statistical significance, the erlotinib–KRAS interaction results must be viewed cautiously, especially since survival results were not overwhelming, the study involved multiple subsets, and was retrospective. Moreover, the relatively small number of patients with KRAS-mutant tumors increases the likelihood of an imbalance in baseline characteristics that could have been responsible for some or all of the observed difference between treatment groups.29
HER2
HER2 amplifications have been observed in lung carcinomas, while HER2 mutations were found to be restricted to the adenocarcinoma histotype.36 The frequency of HER2 mutations was slightly higher in females (4.1%) than in males (1.8%) and in never-smokers (3.1%) than in smokers (1.9%). The mutations show a similar positioning compared with those found in the EGFR gene. Increased copy number of the HER2 gene and overexpression of HER2 are associated with tyrosine kinase inhibitor (TKI) sensitivity in EGFR-positive patients,37,38 possibly because TKIs induce sequestration of HER2 and HER3 receptors in an inactive heterodimer configuration with the EGFR.
MET
The MET gene encodes a transmembrane tyrosine kinase receptor for human growth factor/scatter factor (HGF/SF). Binding of HGF/SF to MET leads to the activation of a number of signaling pathways. MET elicits unique mitogenic and morphogenic effects by stimulating cell–cell detachment, migration, invasiveness, tubule formation, and branching, in addition to proliferative and antiapoptotic activities.39 MET appears to be implicated especially in adenocarcinomas.40 Cigarette smoking induces overexpression of HGF in type II alveolar pneumocytes and lung cancer cells. Overexpression of HGF in lung cancer cells induces alveolar differentiation/proliferation, and MET activation may play special roles in well-differentiated lung adenocarcinomas. Furthermore, HGF seems to have a particular role in the bronchioloalveolar carcinoma (BAC) subtype of pulmonary adenocarcinomas, in which a high level of HGF in broncho-alveolar lavage (BAL) fluid is associated with poorer outcome and is an independent prognostic factor.41 MET amplification showed a trend toward poor prognosis in adenocarcinoma, suggesting that anti-MET could be an alternative therapeutic in this subtype, particularly for EGFR TKI resistance.42
Conclusion
Several layers of evidence indicate that multiple genetic disturbances found in lung adenocarcinomas can be incorporated as predictive markers for metastatic behavior in lung adenocarcinomas. These predictive markers can also be used to identify subgroups of patients who can have a dramatic response when treated with novel targeted therapies.■