Adjuvant systemic treatment of breast cancer is moving away from the limited portfolio of traditional hormonal drugs and chemotherapy, towards a gamma of novel anti-hormonal and chemotherapeutic drugs, immunotherapeutic approaches and small molecules. All these therapeutic approaches are unfortunately effective in a limited number of patients and are often associated with high costs and significant side effects. Therefore, the traditional “one size fits all” approach can no longer be upheld, and a personalised approach based on individual tumour biomarkers is required to arrive at the optimal balance between effectiveness on the one hand and costs and side effects on the other.
Over recent years, progress in molecular techniques has made it possible to analyse formalin fixed paraffin embedded (FFPE) tumour material of larger cohorts of patients. This has incentivised many translational studies relating molecular tumour biomarkers to diagnosis, prognosis and/or response to therapy, yielding many relevant molecular biomarkers that have rather quickly made it to clinical pathology practice. The aim of this paper is to provide an overview of the molecular biomarkers and associated molecular tests that are currently relevant in pathology of invasive breast cancer.
Diagnostic markers
Hereditary breast cancers
About 5–10% of breast cancer cases are due to a hereditary predisposition.1 In most of these cases, mutations are found in well-characterised, medium to high-risk genes, such as BRCA1, BRCA2, CHEK2, TP53, PALB2, or BRIP1,2 of which BRCA1 and BRCA2 are the most important ones. Promotor hypermethylation of BRCA1 and BRCA2 seems to be very infrequent in BRCA1/2 germline mutation related breast cancers, and significantly more frequent in sporadic cancers [data unpublished, submitted for publication]. BRCA1/2 promoter methylation testing, pointing to sporadic cancers when present, may grow out to be clinically useful once the diagnostically optimal CpG islands have been identified.3
Copy number analysis by array comparative genomic hybridisation (CGH) showed frequently occurring gains of 3q, 7p, 8q 10p, 12p, 16p and 17q, and loss of 2q, 3p, 4p, 4q, 5q, 12q, 16p and 18q in BRCA1 germline mutation related cancers. BRCA2 related breast cancers show more frequently gains of 8q, 17q22–q24 and 20q13, and loss of 8p, 6q, 11q and 13q compared to BRCA1 related cancers.4,5 A multiplex ligation-dependent probe amplification (MLPA) kit® (MRC Holland, Amsterdam, The Netherlands) for copy number testing pointing to BRCA1 related cancers is commercially available.
Molecular (intrinsic) typing
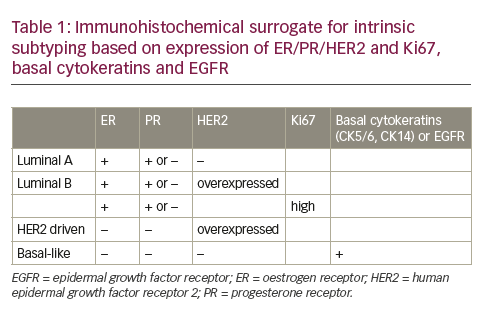
Several gene expression studies have revealed the existence of five molecular subtypes of breast cancer: a “basal-like” subgroup with low oestrogen receptor (ER)/progesterone receptor(PR)/ human epidermal growth factor receptor 2 (HER2) and expression of basal cytokeratins; a subgroup mainly driven by HER2 amplification and overexpression while being ER/PR low; a luminal A group with high ER/PR and low HER2; and a luminal B group that is also ER/PR high but with additional HER2 overexpression and/ or high proliferation.6,7 A further “normal breast like” group is likely the result of absence of tumour in the frozen pieces that were analysed without morphological control by a pathologist. These different classes have varying clinical behaviour and have led to a new way of thinking about classification of breast cancer going beyond morphology. Nevertheless, good correlations exist between these intrinsic subtypes and morphology. The basal-like group contains high grade ductal medullary and metaplastic cancers as well as the low grade salivary gland type cancers (adenoid cystic cancers [AdCC], acinic cell cancers, myoepithelial cancers). The HER2 group contains poorly differentiated HER2 overexpressing ductal and apocrine cancers. The luminal A group contains mainly low grade ductal, lobular, ductulolobular, tubular, cribriform, mucinous and micropapillary cancers. Several intrinsic typing tests are commercially available (PAM50®, NanoString, Washington, US; Blueprint®, Agendia, Amsterdam, The Netherlands). However, these intrinsic subtypes have little added value to type, grade and expression of ER/PR/HER2, and the reproducibility of intrinsic subtyping based on gene expression has proven to be low across datasets and technology platforms. Besides, these tests are not available for local testing in pathology labs, are time consuming and expensive. Therefore, a more practical approach is probably to use an immunohistochemical surrogate as depicted in Table 1.
PAM50 and Blueprint are two commercially available gene expression test for intrinsic subtyping (from centrally at Nanostring and Agendia, respectively). The PAM50 signature employs 50 genes and can be applied on FFPE material.8 Blueprint contains 80 genes and can be applied on FFPE material as well.
Translocations
Adenoid cystic carcinoma
While being a frequent cancer type in the minor and major salivary glands with frequent perineural invasion and poor prognosis, AdCC is a rare cancer type in the breast, accounting for 0.1–1% of breast cancers, with infrequent perineural invasion and indolent clinical behaviour despite their triple negative (ER/PR/HER2) state.9 These carcinomas often display the recurrent chromosomal translocation t(6;9) (q22e23;p23e24), which generates oncogenic fusion transcripts involving the two transcription factor genes MYB and NFIB. In the t(6;9) (q22eq23;p23ep24), the exon 14 of MYB is fused to the final coding exons of NFIB, usually due to breakpoints in MYB intron 14 and intron 8 in NFIB. The fusion results in loss of the 3’-end of MYB, including several conserved binding sites for microRNAs that regulate MYB expression negatively. More recently, also recurring t(8;9) and t(8;14) translocations have been described fusing the MYBL1 gene to the NFIB and RAD51B genes, respectively.10,11 Due to the characteristic histological features of AdCC, translocation assays may not be often necessary in diagnostic practice, but may be assessed by fluorescent in situ hybridisation (FISH) kits or reverse transcription polymerase chain reaction (RT-PCR) in difficult cases.
Secretory cancer
Secretory cancer is a rare breast cancer type that occurs at all ages, even at quite young age. Recently, it was shown that secretory cancer is characterised by a balanced chromosomal translocation t(12;15) (p13;q25), which leads to the formation of an oncogenic ETV6-NTRK3 fusion gene encoding a chimeric tyrosine kinase, also demonstrated in paediatric mesenchymal cancers.12 Translocations may be assessed by FISH or RT-PCR in difficult cases.
Prognostic markers
Gene expression tests
Several gene expression tests have been developed to predict prognosis and response to therapy. The above described PAM50 has proven prognostic value.8 Further popular prognostic gene expression tests are EndoPredict® (Sividon Diagnostics, Köln, Germany), MammaPrint® (Agendia, Amsterdam, The Netherlands) and Oncotype Dx® (Genomic Health, California, US).
EndoPredict is an 8-gene RT-PCR test that primarily targets ER+/HER2- patients that have 0-3 lymph node metastases. It can be applied on FFPE and has been validated for local application in pathology labs, and is available from Sividon Diagnostics. It was proven to be prognostically important in several studies (level I-II evidence), especially in combination with lymph node status and tumour size,13 and gives a binary prognostic indication. Mammaprint is a 70-gene microarray expression test that primarily targets stage I-II breast cancer patients with tumour size <5 cm. It was originally developed as a prognostic factor with evidence level I on frozen material but was recently also validated on FFPE material.14 It is available only centrally from Agendia and gives a binary prognostic indication. Oncotype Dx is a 21 gene RT-PCR test (including five reference genes) targeting ER+ patients with no lymph node metastases. It is available only centrally from Genomic Health and gives a three-tiered prognostic indication.15
These gene expression tests in general provide good prognostic information and are analytically well validated except. A recent paper indicated that these tests may be significantly suffer from intra-tumour heterogeneity.16 They are recommended in most guidelines like AGO, St Gallen, The European Society for Medical Oncology (ESMO), The American Society of Clinical Oncology (ASCO), National Comprehensive Cancer Network (NCCN), and The National Institute for Health and Care Excellence (NICE). A hurdle in broader application is the high costs at about €3000 per test. The added prognostic value to type and grade is likely limited,17 but their better reproducibility are arguments in favour. No prognostic comparative studies have been performed, so decisions on the preferred test are usually taken based on local arguments.
IHC4
Although the immunohistochemistry (IHC)4 score is based on IHCbased protein expression of ERα, PR, HER2 and Ki67 and thereby not strictly a molecular test, it is discussed here since it has been proposed as a cheap alternative to gene expression tests. IHC4 provided similar prognostic information as Oncotype Dx score in a study of ER-positive patients from the ATAC (Arimidex, Tamoxifen, Alone or in Combination) trial who did not receive adjuvant chemotherapy.18 In addition, PAM50 and IHC4 scores provided comparable prognostic information in the HER2-negative/node-negative group.19 Quantitative evaluation of ER, PR and HER2 status is well standardised and guidelines for Ki67 assessment are available.20 IHC4 could significantly improve the routine availability of prognostic information in a cost-effective manner.
Predictive markers
Gene expression tests
The value of the above described gene expression tests EndoPredict, MammaPrint and Oncotype Dx lies in their predictive value rather than their prognostic power in view of their high costs. If patients that are deemed eligible for chemotherapy based on classic prognostic factors can be safely spared chemotherapy when their gene expression test turns out to be favourable, much money can be spared elsewhere and application may be cost-effective. All these tests have proven to have such predictive value at level I evidence, although lack of intra-tumour heterogeneity studies remains a weak point. Further, comparative predictive studies have been performed, so decisions on the preferred gene expression test are usually taken based on local preferences rather than firm scientific data.
ERα mRNA expression
Since IHC has its limitations, ESR1 mRNA expression has been explored as an alternative. There appeared to be a very good correlation between ESR1 mRNA expression by microarray analysis and ERα IHC.21 There is no proven predictive superiority of gene expression in mRNA-protein discrepant cases. The “Targetprint” test (Agendia, Amsterdam, The Netherlands) was recently taken off the market.
ESR1 mutations
ERα expression has for a long time served as indication for hormonal therapy. More recently, molecular changes in ESR1, the gene on chromosome 6 encoding ERα, have been demonstrated to relate to endocrine responsiveness as well. ESR1 mutations leads to conformation change which mimics activated ligand-bound receptor and induces ligand-independent ER activity,22 resulting in tumour growth despite endocrine therapy. The mutants appear to retain some sensitivity to drugs directly targeting the receptor suggesting potential pharmacologic strategies for these patients.23
Although ESR1 mutations are very uncommon in primary breast tumours,24 metastases more frequently harbour mutations in the ligandbinding domain resulting in endocrine resistance.25,26 Interestingly, these mutations have been demonstrated in blood providing a monitoring option.27 Breast tumours of women naïve for endocrine therapy do not seem to harbour these mutations, suggesting either the clonal selection of very rare resistant primary tumour clones or their later acquisition under the pressure of endocrine treatments.25 Relatively many patients develop polyclonal mutations. Aromatase inhibitor treatment in the metastatic setting rather than in the adjuvant setting induces ESR1 mutations.28 The pan-phosphatidylinositol 3-kinase (PI3K) inhibitor pictilisib and the CDK4/6-inhibitor palbociclib do not seem to be effective in ESR1 mutated patients,29 and ESR1 mutations did not indicate impaired survival on fulvestrant compared to ESR1 wildtype. On examestane, however, ESR1 mutations do seem to indicate worse prognosis. Overall, ESR1 mutations seem to be associated with more aggressive disease biology.29
ESR1 amplification
The frequency of ESR1 amplification is highly debated. ESR1 gains and amplifications were originally described in 15% and 20% of breast cancer samples, respectively.30 In these studies, ESR1 copy number increase was correlated to high ERα protein expression, and was associated with very good response to tamoxifen and good survival. In later studies, the ESR1- amplification frequency was 0–23% when FISH was used for detection, and less than 5% when biochemical assays, CGH, MLPA, and qPCR, were used.31 In general, the amplification frequency is highly concordant between the different studies that used biochemical assays, whereas considerable variability exists among the studies that used FISH (different protocols, scoring systems, artefacts). Most amplifications are low level, and FISH may be hampered by presence of pre-mRNA.31
ESR1 promoter hypermethylation
ESR1 promoter methylation has been suggested as an alternative mechanism of ERα expression loss. Methylation of the ESR1 promoter is found in a proportion of the ER negative tumours.32 and in ER negative cells lines. The use of demethylating agents was shown to reactivate ER expression which reveals a potential new strategy to overcome endocrine resistance in breast cancer patients.33 ESR1 promoter methylation analysis has however not yet reached the status of a clinical test.
HER2 amplification
The HER2 is a proto-oncogene amplified in 10–15% of human breast cancers. HER-2/neu (HER2) status predicts prognosis and sensitivity to HER2 protein targeting drugs such as trastuzumab, pertuzumab and lapatinib. Accurate assessment of HER2-status for breast cancer patients is therefore crucial, which can be done by IHC, FISH, chromogenic in situ hybridisation (CISH), silver in situ hybridisation (SISH) and MLPA, as described by Moelans et al.34 IHC is the most widely used since it is cheap, fast and simple. Staining is usually scored according to the DAKO guidelines as 0 (no staining or <10% of the tumour cells are weak and incomplete, not circumferential), 1+ (≥10% of the tumour cells is weak and incompletely stained), 2+ (≥10% of the tumour cells are stained moderately intense [complete, or incomplete], or 10% is stained strongly and circumferentially).35 However, membrane staining can be influenced by fixation conditions, and IHC may miss amplified cases.36 FISH is an effective and sensitive method to assess HER2 gene amplification status. However, it not a very practical primary screening tool for HER2 copy number status since it is time consuming, signals fades over time, tumour morphology is more difficult to appreciate and tumour heterogeneity can easily be missed. FISH is therefore usually performed as a second-line test for the IHC (2+) equivocal group. The morphology with CISH and SISH is much better preserved, slides can be kept permanently, and it is easier to score. A recent review showed that HER2 IHC, FISH, CISH and SISH have good concordance between them.34
MLPA is a multiplex PCR technique that has been well validated for assessment of HER2 copy number status that works on only small amounts of fragmented DNA isolated from FFPE material (50–100 ng). MLPA can analyse up to 45 genes in one assay, and is thus able to simultaneously test different genes that are important for therapy selection or prognosis. Different studies have shown a good correlation between MLPA and IHC, FISH and CISH,36 but MLPA has not been directly validated with regard to its predictive value for trastuzumab response.
All these techniques show an overall good correlation with each other in comparative studies, but not without discrepancies both ways.34 As a result, there is no gold standard HER2 test, and no consensus on which of the above-mentioned techniques is to be preferred. A clinical diagnostic test for HER2 should accurately determine HER2 protein levels or gene copy number and should be reproducible and precise across multiple sites and users. Decisions on how to test can therefore be taken locally after proper local validation. Local preferences may also determine whether to use amplification tests only as reflex test in cases that are equivocal by IHC (2+), or as primary screening tests, or parallel next to IHC.
Traditionally, polysomy of chromosome 17, where HER2 is located, has received much focus as a source of “pseudo-amplification” of HER2. Therefore, many FISH and CISH kits have dual probes for HER2 and a centromere 17 sequence, usually CEP17, to enable calculation of a HER2/CEP17 ratio, which according to guidelines, needs to be above ≥2.0 to indicate “true” HER2 amplification.37 However, several studies have shown that polysomy of chromosome 17 is virtually absent in breast cancer,38 but that 17q rather shows a very complex gains and losses pattern, unrelated to the copy number status of the centromere. Correction with CEP17 probes may thereby provide misleading HER2 gene status assessment results, and the absolute copy number of HER2 likely serves best to indicate eligibility for trastuzumab therapy.39
HER2 mutations
In breast cancer, the potential therapeutic opportunities offered by rare (1%-3%) HER2 somatic mutations were long neglected, given the high prevalence of HER2 gene amplification.40 The development of massive parallel sequencing has shed new light on these activating mutations, mostly present in the tyrosine kinase domain (68%, exons 19–20) and the extracellular domain (20%, exon 8).41 Tumours considered to be HER2- negative based on current guidelines may still be “addicted” to HER2 signalling due to HER2 activating mutations and hence may also benefit from HER2 targeting agents. Bose et al. have functionally characterised 13 HER2 mutations using in vitro kinase assays, protein structure analysis, cell culture and xenograft experiments.41 Seven of these mutations were activating mutations and all of these mutations were sensitive to the irreversible HER2/EGFR tyrosine kinase inhibitor neratinib, thus validating HER2 somatic mutations as drug targets for breast cancer treatment.41 This finding may very well lead to a new era for breast cancer management, where HER2 status is no longer solely determined by IHC and ISH techniques. However, clinical trials are first needed to test whether HER2- mutated tumours are responsive to HER2-targeted drugs, and if so, which mutations are predictive of sensitivity to which agent.
Gene expression arrays for HER2
HER2 mRNA expression has also been explored as an alternative predictive test. There was a very good correlation between HER2 mRNA expression by microarray analysis and HER2 IHC.21 There is proven predictive superiority of gene expression in discrepant cases. The “Targetprint” test was recently taken off the market. Oncotype Dx also contains the HER2 gene.
PIK3CA mutations
PIK3CA is an oncogene exhibiting oncogenic gain-of-function mutations in several cancers occurring in 20–40% (25% by COSMIC; July 2013) of breast cancers. Several studies have suggested that PI3K pathway activation can negatively influence response to trastuzumab therapy,42,43 and endocrine therapy.44
Given their frequency, oncogenic capabilities, and the potential to induce resistance to commonly prescribed breast cancer treatments, the clinical relevance of PIK3CA mutations needs to be further clarified. In light of the emergence of a broadening array of anti-HER2 agents, determination of PIK3CA mutational status could have important clinical utility. In addition, inhibitors of the PI3K pathway that may reverse acquired and de novo drug resistance are currently in clinical development.
BRCAness
BRCAness refers to a tumour genotype as seen in BRCA germline related cancers. Such a phenotype predicts response to high dose chemotherapy45 and poly ADP ribose polymerase (PARP) inhibitors. Although yet far from completely defined, several techniques may be employed to identify BRCAness. Copy number profiles by array CGH or MLPA46 that point to BRCA germline mutation related cancers as described above may be useful. Further, BRCA1/2 promoter methylation may lead to a similar phenotype by inactivating BRCA1/2 function.47 Finally, tumours may be sequenced for sporadic BRCA1/2 mutations. This is expected to become more and more clinically important in breast cancer treatment.
Testing of metastases
Despite advances in treatment and screening modalities, approximately 30% of breast cancer patients eventually develop recurrent or metastatic disease. Although metastatic breast cancer remains essentially incurable and therefore the main cause of breast cancer related death, it is a highly understudied field. Growing evidence describes extensive differences in ER, PR and HER2 protein expression between primary breast tumours and their paired metastases (termed conversion).48 Also, gene expression tests of these markers are worth executing, since the robustness of these tests can lead to a specific insight of the true extent of conversion. This can have an impact on systemic treatment choice and efficacy.
As mentioned earlier, many predictive and prognostic tests are already in use for primary tumours. Since metastases are a reflection of poor prognostic disease, especially predictive tests could be valuable to also test in this group. However, if described gene expression tests as Endopredict, MammaPrint and Oncotype Dx are equally able to foretell response to therapy in metastases, remains to be elucidated.
Next generation sequencing (NGS) of ‘cancer-associated’ genes already revealed high genomic concordance between breast tumours and paired metastases [data unpublished, submitted for publication], but new mutational events emerge during progression. Although not present in every patient, this clonal evolution can lead to new targetable aberrations in metastases relative to their tumour of origin.49 Currently, the above described potentially targetable ESR1 ligand-binding domain mutations occur especially in metastases and testing for this is probably the most established molecular test on breast cancer metastases.
Some of the differences between primary breast tumours and metastases can be explained by evolutionary pressure of systemic therapy. An example can be found in the PI3K/AKT/mTOR pathway, since adjuvant endocrine therapy can significantly increase p-mTOR, p-4EBP1 and p-p70S6K expression in metastases, indicating the compensatory activation of this pathway.50 Since this finding could forebode acquired endocrine therapy resistance,51 testing of metastases could prevent maladjusted treatment.
Recapitulating, molecular diagnostics could disclose new, (potentially) druggable targets in metastases compared to the primary tumour and could be extremely valuable for personalised cancer treatment in metastatic breast cancer.
The role of next generation sequencing
NGS allows for the simultaneous mutation analysis of either the genome, exome, or genes of interest. For routine diagnostics targeted NGS is a reliable assay to detect (hotspot) mutations in FFPE
samples.52 Although this technique is already extensively used for other tumour types (e.g., lung and colon tumours), mutation analysis of breast cancer is not yet routine practice. Implementation of NGS for breast cancer specifically analysing mutations in HER2, ER and PIK3CA, would provide information on treatment options. Potentially, BRCA mutation analysis can be added in the near future, if clinical studies show that the presence of a BRCA mutation is indication for treatment with PARP inhibitors.53
Next to mutation analysis, NGS allows for simultaneous copy number variation (CNV) detection,54 so HER2 amplification analysis can be performed with the same NGS assay as mutation analysis. For reliable CNV detection the percentage of neoplastic cells in the sample needs to be preferably at least 30%. Since NGS provides a combined result of all normal and neoplastic cells within the tissue, the tumor information is diluted which might result in a negative CNV result when too few neoplastic cells are present in the sample.
NGS can in principle also be used for translocation and methylation detection in FFPE tissue. Currently, mostly genome wide RNA-seq and bisulfite sequencing approaches are used, which are not suitable for fast routine diagnostics. However, amplicon based NGS panels can enrich cDNA samples for fusions. Also, novel fusions, where only one of both fusion partners is known, can be detected using this technique. For methylation analysis, currently relatively large amounts of DNA are required due to the bisulfite treatment which degrades DNA, so this is not a technique which can easily be implemented in routine diagnostics. Also for methylation detection, a targeted NGS approach can be used, where the primers should be designed for bisulfite treated DNA. As both translocation and methylation analysis require a different starting material compared to mutation and CNV analysis (RNA and bisulfite treated DNA compared to untreated DNA), combining all techniques in one assay will only be possible for the sequencing step and not the preceding library preparation. In summary, NGS can currently be used to detect mutations and CNV in FFPE samples using a targeted approach.
Liquid biopsy
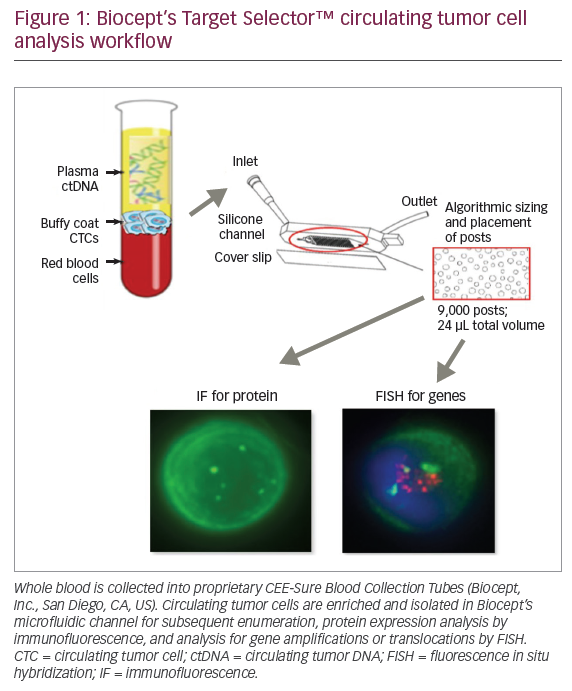
Although the above mentioned techniques can be performed reliably on tumour specimens and small biopsies, the question remains whether less invasive methods can be applied. Especially for the follow-up of patients to evaluate response to therapy, liquid biopsies seem to be an excellent alternative to tissue biopsies. Therefore, the tumour first needs to be screened for specific molecular alterations, which can subsequently be followed in the liquid biopsy, for example in blood samples. From the blood, either circulating tumour cells (CTCs) or circulating free tumour DNA (ctDNA) can be extracted to study these molecular alterations. Since the amount of CTCs or ctDNA in the blood is limited, very sensitive techniques are necessary, such as a digital PCR. This technique allows the detection of ctDNA mutations with a frequency of as low as 0.1%. It is therefore also possible to detect minimal residual disease in blood samples.55 Alternatively, NGS can be used when a high coverage is used to allow very sensitive mutation detection. Rothé et al. described that known mutations can be detected in the blood using a targeted NGS approach when a coverage of 25,000x is used.56 Since the isolation of CTCs is more challenging than the isolation of ctDNA, clinical studies are ongoing to test whether CTCs can be used for monitoring treatment response.56,57 In conclusion, liquid biopsies seem a reliable, non-invasive method to detect low-level molecular alterations for the follow-up of cancer treatment.
Conclusions
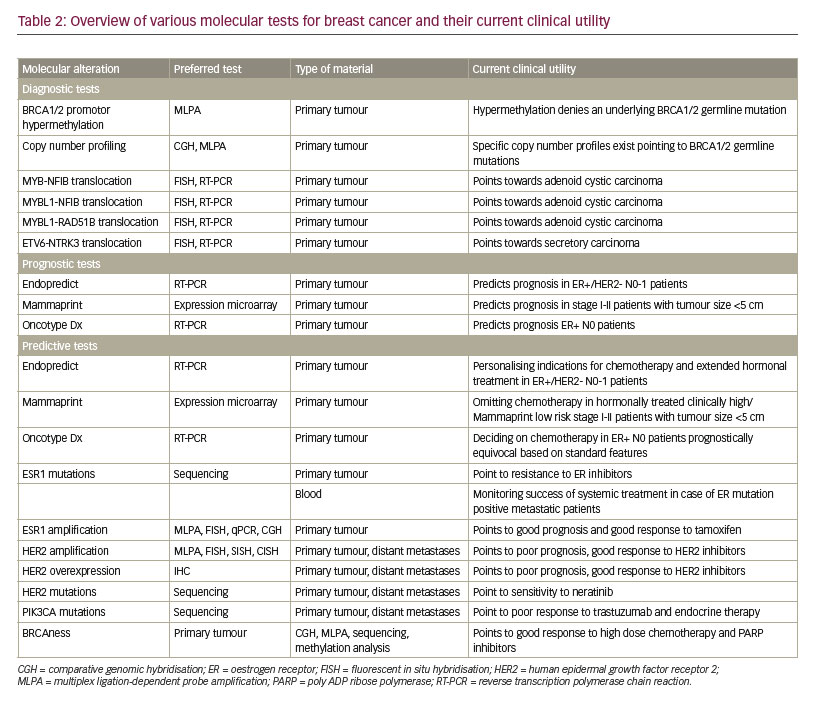
The portfolio of diagnostic molecular tests for breast cancer is constantly expanding. Tests that have been established for routine use include BRCA1/2 profile testing for detection of hereditary cancers by copy number MLPA/NGS and MS-MLPA, translocation testing for AdCC and secretory cancer, predictive gene expression profiling in clinically equivocal cases, HER2 amplification testing by ISH/MLPA, ERα mutation testing in metastases, HER2 and PIK3CA mutation testing in case of trastuzumab treatment, and BRCAness testing by MLPA (hypermethylation, copy number) or NGS (copy number, mutations). Table 2 provides an overview of the currently useful molecular methods in breast cancers, their preferred way of testing and the clinical utility.