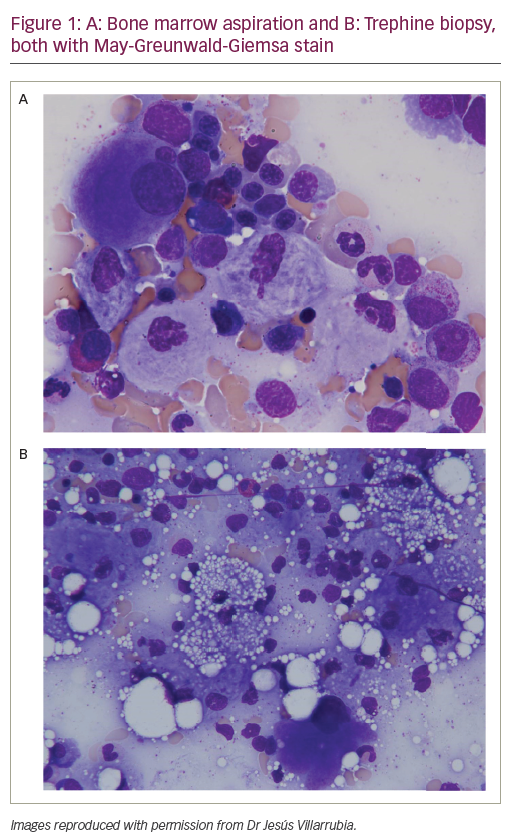
Venomous snakebites are still a public health problem in tropical countries. Even in metropolitan areas, such as Bangkok, green pit viper (Cryptelytrops albolabris and C. macrops) bites are still prevalent. As we are invading their territory, they adapt to live among humans in our backyards. With some exceptions, clinical syndromes of snakebites can be divided simply into two classes: elapid envenomation attacks neuromuscular junctions, causing muscular paralysis; viper envenomation targets the haemostatic system, resulting in bleeding disorders. Therefore, viper venoms are of great interest to haematologists. Vipers (Viperinae) are members of the snake family that develops the most advanced venom-injecting organs, the long sheathed fangs that can be folded like switchblades. Some vipers possess pit organs, infrared detectors for locating warm-blooded prey in the dark. This specialised subfamily is termed crotalidae (Pit viper), separating them from viperidae (True viper). Vipers found in the Americas are pit vipers, while African vipers are true vipers. Asia and Europe are homes to both subfamilies.
Viper Bites
Clinically, viper bites cause both local tissue damage and systemic effects: local tissue swelling, blisters from dermo-epidermal separation, ecchymoses and gangrene. Snake venom metalloprotease (SVMPs) have been implicated in these tissue injuries, both directly and indirectly via vascular damage1 and inflammatory responses.2 The most prominent systemic effects of pit viper venoms are towards fibrinogen and platelets. Although the venoms clot fibrinogen (thrombin-like effect) and activate platelets in vitro, these friable clots and platelet aggregates are rapidly cleared from circulation in vivo, resulting in hypofibrinogenaemia, termed defibrination syndrome, and thrombocytopenia.3 On the other hand, true vipers, such as Russell’s viper and Daboia russelii, activate common pathways of coagulation via factor X and V, causing bleeding from disseminated intravascular coagulation. Kinetics studies in humans showed that the viper venom is rapidly absorbed. However, the effects on blood may be delayed and prolonged as the half-life of viper venoms is over 24 hours.4 Antivenoms, the specific treatment of venomous snakebites, are polyclonal antibodies derived from horses or sheep. Older versions of antivenoms may cause potentially fatal early reactions mediated by complement activation from the immunoglobulin (Ig)-G (fragment, crystallisable) Fc portion.5,6 Current F (ab)’2 antivenoms in Thailand, the products of Queen Saovabha Memorial Institute, are pure and devoid of the Fc portion. A study revealed that the incidence of reactions is low (3.5%) and unpredictable by the intradermal test.7 Therefore, a hypersensitivity skin test for immunoglobulin (Ig)-E-mediated reactions is unnecessary, but close observation is essential during administration in all patients. Although antivenoms are efficacious in reversing coagulopathy and thrombocytopenia from viper venoms, this efficacy in treating local tissue injuries is limited.8,9 This is consistent with animal studies showing that venom-induced local damages was rapid, and unable to be treated even by immediate antivenom.10 Therefore, local tissue injury is the current management problem after snakebites, and further research is needed.
Viper Venoms
The significance of viper venoms goes far beyond the tropical medicine discipline. Different snake venoms have been found to affect almost all components of the haemostatic system, including the vascular wall, platelets, clotting factors, clotting inhibitors and fibrinolytic components. In addition, there are effects on a wide variety of physiological and pathological processes, including angiogenesis, inflammation and cancer metastasis. Consequently, snake venoms are rich sources of proteins with potential to be useful diagnostic or therapeutic agents or compounds for studying the molecular mechanisms of haemostasis. Apart from the conventional protein purifications from snake venoms, current molecular biology permits the construction of a complementary DNA (cDNA) library of the expressed genes in venom glands. This provides us with complete and accurate sequences of DNA and conceptually translated proteins to correlate with their functions. Furthermore, cDNA is an unlimited source of pure recombinant proteins that can be used to investigate their therapeutic and/or diagnostic values. Finally, mutagenesis can be performed to study the structure–function relationship and engineer proteins with desirable activities.
The evolutionary origins of venom proteins are from non-toxic genes that are recruited to express in venom glands, the special organs that started developing before snakes phylogenetically branched out from lizards.11 For example, snake venom serine proteases (SVSPs) are from tissue kallikrein, SVMPS/disintegrins are from a disintegrin and metalloproteinase (ADAM) proteins and a prothrombin activator, is from clotting factor X.12 Subsequently, these genes acquired toxic properties following a process known as accelerated evolution.13 After gene duplications and high rates of non-synonymous base substitutions in coding regions, a wide variety of novel functions emerged, greatly expanding the arrays of susceptible prey with enormous geographical and seasonal variations.
Snake venom proteins, like other toxins, are cysteine-rich (CR) peptides.14 In an oxidising extracellular environment most form disulphide bonds. The conserved bonds may be helpful in supporting the original folding structures of proteins during accelerated evolution. On the other hand, alterations of disulphide bond patterns may lead to novel structures and functions enhancing their variety. For example, a loss in a disulphide bond in the disintegrin-like domain of SVMPS resulted in a disintegrin, the integrin-blocking proteins unique to snake venoms.15 In addition, changes in disulphide bonds may cause different forms of post-translational modifications, such as the release of disintegrin domains from the protease domain of SVMPs.15 Furthermore, disulphide bonds can determine the multimeric structures, e.g. dimer versus tetramer, of the venom CLPs. Only the tetramers can cluster platelet receptors, resulting in stronger platelet activation.16 Disulphide bonds also linked two peptides, either in the same family, e.g. a dimeric disintegrin, or different families of proteins, e.g. SVMP and CLPs. These create novel and/or composite functions. Although cysteine bonds may be beneficial for snake venom evolution, they make these proteins difficult to recombinantly express, even in eukaryotic systems.
Snake Venom Serine Proteases
Almost all of the enzymes in coagulation cascade and fibrinolysis are serine proteases characterised by the catalytic triad: serine (S), aspartate (D) and histidine (H). However, most SVSPs are originate from the glandular kallikrein that later evolved to acquire coagulation and fibrinolytic/fibrinogenolytic activities. This may explain hypotensive effects of some enzymes due to the kallikrin-like activity producing kinin. The largest group of venom enzymes in this family is termed ‘thrombin-like’ enzymes due to their activities, not the evolutionary origin.
Thrombin plays pivotal roles in haemostasis.17 The inactive precursor, prothrombin, is composed of the N-terminal domain that is cleaved from the C-terminal catalytic domain or thrombin upon activation. In contrast to factors VII, IX, X, which require their N-terminal domains and co-factors (tissue factor, factor VIII and factor V, respectively) to form proteinase complexes on membrane, thrombin is released freely in solution to perform its myriad functions. The primary function of thrombin is the conversion of fibrinogen to fibrin by cleaving fibrinopeptide A and, subsequently, fibrinopeptide B from Aα and Bβ chains of fibrinogen, respectively. On the fibrin surface, thrombin activates factor XIII, resulting in fibrin cross-link. It also amplifies the positive feedback loops of coagulation by activating factors V, VIII and XI, as well as platelet protease-activated receptor (PAR). The activation of PAR not only stimulates platelet aggregation, but also provides a negatively charged surface, supporting blood coagulation. The activation of factor XI and PAR is dependent on the ability of thrombin to bind platelet glycoprotein (GP) Ib. Furthermore, in the presence of the co-factor thrombomodulin (TM), thrombin activates protein C and thrombin-activatable fibrinolysis inhibitor (TAFI) to inhibit coagulation (via degradation of factors Va and VIIIa) and fibrinolysis (via digestion of plasminogen activator-binding sites on fibrin), respectively. Molecular mechanisms of the multifunctionality of thrombin have been under extensive investigation. Apart from the active site of thrombin, there are mixed positive charges and hydrophobic patches on both sides of the molecules known as anion-binding exosites I and II (ABE I and ABE II) that confer thrombin substrate specificity (see Figure 1). ABE I is responsible for fibrinogen, factors V, VIII, XI, PAR and TM contacts, while the ABE II binds platelet GPIb and chondroitin sulphate part of TM.18 Thrombin inhibitors may target the ABE I (hirudin) exosite II (heparin and hemadin) or both exosites (Bothrojaracin, a C-type lectin from the snake Bothrops jararaca).
Thrombin-like enzymes are named according to the ability of venom proteases to clot fibrinogen in vitro. Notably, these enzymes do not contain the full effects of thrombin. They most commonly release only fibrinopeptide A, termed venombin A, for example ancrod from the Malayan pit viper, Calloselasma rhodostoma,19 batroxobin from B. atrox20 and GPV-TLs from the green pit viper, C. albolabris;21 or only fibrinopeptide B, termed venombin B, for example contortrixobin from Agkistrodon contortrix,22 or both fibrinopeptides, termed venombin AB, for example bilineobin from A. bilineatus.23 However, bilineobin preferentially cleaves fibrinopeptide B. In addition, factor XIII is not activated, resulting in friable clots subjected to fibrinolysis. Some SVSPs show other activities of thrombin. For example, platelet PAR is activated by PA-BJ from B. jararaca.24 A TM-independent protein C activator, ACCC, was isolated from A. contortrix contortrix,25 and a factor V activator, RVV-V, was purified from Russell’s viper, D. russelii.26 Other SVSPs contain fibrinogenolytic27 and/or plasminogen-activating activities.28
Sequence analysis of thrombin-like SVSPs could not define ABE I and II homologous to thrombin.21 This is not surprising because their functions do not have the same multiplicity and their evolutionary origin is different from that of thrombin. However, parts of molecules, which may be called ‘exosites’, away from the active site are likely to play roles in substrate recognition. Unfortunately, there is no reported high-resolution crystal structure of a fibrinogen-clotting enzyme; only crystallography data of plasminogen activator28 and protein C activator29 are available. Interestingly, the overall structures of these two are similar. Structure– function correlations of SVSPs require further study.
The properties of thrombin-like enzymes have been employed for diagnostic and investigational uses. For example, venombin A and venombin B were used to define the contributions of fibrinopeptide A or B releases in fibrin polymerisation.30 In addition, SVSP from B. atrox (reptilase time) can determine the causes of prolonged thrombin time because the test is unaffected by heparin. The protein C and factor V activators are used for activated protein C resistance and lupus anticoagulant tests, respectively. Furthermore, thrombin-like enzymes have the potential to be therapeutic anticoagulants.31 However, a large randomised trial revealed that ancrod is not better than placebo in the treatment of ischaemic stroke.32
Snake Venom Metalloproteases/Disintegrin
SVMPs are zinc-dependent multidomain proteins. The nascent peptides contain a signal peptide and a pro-sequence that keeps the enzyme inactive. The mature secreted proteins from class P-I comprise only a metalloproteinase domain, while class P-II contains additional disintegrins and P-III class possesses disintegrin-like and CR domains. The disintegrin domains are usually released from the mature proteins. Free disintegrins interfere with cell–matrix interaction via integrin binding, while attached disintegrin-like or disintegrin domains target the enzymes to the sites of actions.33 Several variants of these domain structures have been reported, including attached disintegrin domains, free disintegrin-like domains, homo- or heterodimers and additional heterodimeric CLPs that joined the C-terminus of proteins by disulphide bonds. The prototype of this class is RVV-X, a factor X activator from Russell’s viper.
Apart from tissue damage, some SVMPs can degrade fibrinogen/fibrin.34 Others digest platelet membrane GPs and their ligands, e.g. GPIb, von Willebrand factor (vWF), collagen and P-selectin GP ligand-1 (PSGL-1). Interestingly, alborrhagin from C. albolabris35 activates the collagen receptor GPVI, causing platelet activation. Furthermore, the SVMPs with lectin subunits can activate prothrombin (Ecarin from saw-scaled viper, Echis carinatus)36 or factor X (RVV-X from D. Russelii). The crystal structure showed that the lectin domain bound the γ-carboxyglutamic acid (Gla) domain of factor X facilitating the action.37
The coagulation factor activators have been used as the diagnostic tools. The prothrombin activator is used to monitor thrombin inhibitor therapy (Ecarin time) and factor X activator to assay for lupus anticoagulant. Furthermore, the modified fibrinogenolytic enzyme alfimeprase has been used successfully as a direct thrombolytic agent for peripheral arterial occlusion in an animal model.38 The advantage of this agent is its rapid onset of action and neutralisability by plasma α2 macroglobulin.39 Unfortunately, phase III clinical trials failed to prove its benefit over standard treatment for acute arterial thrombosis or catheter occlusion.
Disintegrins are venom-derived low-molecular-weight peptides that bind and block integrins. Integrins are cell-surface proteins mediating interactions between cells and extracellular matrices or other cells. They are essential for cell survival, proliferation, differentiation and/or activation. Integrins comprise two transmembrane subunits, α and β. Permutations of the two create a variety of receptors with distinct ligand specificities and functions. The most important integrin in haematology is the platelet αIIBβ3 (GPIIb/IIIa), the major fibrinogen receptor-mediating platelet aggregation. The main feature of disintegrins is the tri-peptide motif that binds integrins including the archetypical RGD or KGD or other variants, WGD, VGD, MGD, MLD, RTS and KTS. The sequences of and around this motif, as well as of the C-terminal tails, determine the integrin-binding specificity.40–42 Although data suggested that KGD conferred disintegrin specificity to GPIIb/IIIa, we found that the KGD-containing albolatin disintegrin inhibited only collagen-induced, not ADP-induced, platelet aggregation.43 The structures of disintegrins showed the integrin-binding motif on the tip of a loop with the C-terminal tail nearby (see Figure 2).44 In contrast to integrins, disintegrin-like domains do not contain this motif and usually attach to a CR domain. Crystallographic data disclose that the conformations of disintegrin-like domains are totally different from disintegrins and the XCD motif in the proteins do not bind integrins, as previously hypothesised.45 Notably, the disintrigin-like and CR domains form a C-shaped structure connecting to the terminal hypervariable region (HVR) that binds specifically to the substrates (see Figure 3).
The most prominent action of RGD, KGD or WGD disintegrins is platelet inhibition. Furthermore, some disintegrins contain anticancer activities by inhibiting integrin-dependent tumour cell migration, metastasis and angiogenesis.46 For example, rhodostomin from C. rhodostoma, trigramin from Trimeresurus flavoviridis and albolabrin from C. albolabris can inhibit cancer cell invasion in an animal model.47 Contortrostatin, a homodimeric RGD-disintegrin from A. contortrix contortrix, induces vascular apoptosis and inhibits angiogenesis.48
Eptifibatide, a derivative of the disintegrin barbourin, is now available as an antiplatelet for coronary diseases.49 In addition, contortrostatin was shown to block breast cancer growth in a mouse model. The preparation was packed in liposome to evade immune system and platelets and to increase half-life.50
C-type Lectin-like Proteins
CLPs are evolved from a large family of the C-type lectins that widely express from microbes to humans.51 However, CLP are neither C-type (calcium-dependent) nor lectin (carbohydrate-binding) proteins.52 The basic building block of CLPs is a heterodimer of a subunit a (α chain) of 14–15kDa and a subunit b (β chain) of 13–14kDa linked by inter-chain disulphide bond. CLP complexes may be αβ, (αβ)2, (αβ)4 or α1α2β1β2.53–55 X-ray crystallography revealed the domain swap between an αβ dimer providing a concave surface for binding to various target proteins.56 The structure of higher multimer of αβ dimer has also been solved (see Figure 4).57
Anticoagulant CLPs bind the Gla domain of clotting factor IX and/or X in the presence of Ca2+.58 Other targets of CLPs are platelet GPIb and/or GPVI or vWF. The GPIb-binding CLPs can cause either platelet agglutination or inhibition in vitro depending on the stoichiometry of bindings.59 However, the affinity of binding is probably lower than vWF, resulting in competitive inhibition in vivo. Some CLPs are disulphide-linked higher multimers and bind GPVI, with or without GPIb-activating signal transduction in platelets. Higher multimer has been implicated in receptor clustering and, thus, activation.16 Examples are alboaggregin A (α1α2β1β2) and alboluxin ((αβ)3), which activate GPVI and GPIb,60,61 while convulxin (αβ)4 bind GPVI and GPIa/IIa, another collagen receptor. In contrast, stejnulxin62 and trowaglerix can activate only GPVI, whereby trowaglerix also causes subsequent GPVI shedding and later platelet inhibition.63 Furthermore, dimeric botrocetin and bitiscetin target vWF and/or GPIb, enhancing their interaction.64 Finally, bothrojaracin, the (pro)thrombin exosite binder, demonstrates a novel mechanism of anticoagulation.
Platelet-targeting CLPs are used to dissect the mechanisms of platelet activation mediated by each GP receptor. In addition, bothrojaracin could treat a rat model of thrombosis.65
Phospholipases A2
The enzymes originating from extracellular phospholipase A2 (PLA2) are animal enzymes that help gastrointestinal digestion and mediate inflammatory and antimicrobial activities.66 The anticoagulant properties have been reported in snake venom PLA2 and are independent of phospholipase activity. The mechanism is likely to be direct binding, hence inhibiting the activated clotting factor Xa or prothrombinase complex.67 PLA2 that binds vascular endothelial growth factor (VEGF) receptor, promoting angiogenesis, has been reported.68
Perspectives
Snake-venom proteins are proven useful tools for diagnosis and investigation as they affect almost all aspects of the haemostatic system. However, therapeutic applications of venom derivatives are limited by potential immunogenicity and the parenteral route of administration, making them unsuitable for repeated treatments. In addition, they have to compete with a lot of efficient and safe antithrombotics and anticancer treatments on the market or in development. Identification of diseases or patient subgroups without effective remedies, employment of venom components to study pathogenesis and, then, production of small molecular mimetics as treatments may be the way of drug discovery in the future. ■